Combining machine learning and dynamic system techniques to early detection of respiratory outbreaks in routinely collected primary healthcare records | BMC Medical Research Methodology

The main steps in MMAING’s workflow, from data pre-processing to outputs, are illustrated in Fig. 1.
In the pre-processing stage, the time series of URTI-related PHC data are obtained, consolidating an input database (green box in Fig. 1). The database is split into training and validation data sets as usual in unsupervised ML protocols (orange and blue boxes). At the same time, a full database is used to implement the NGM. The training datasets are also used to generate synthetic series to enable quantitative evaluation of the models through statistical metrics.
Outbreak detection and early warning signals (EWS) issuance by MMAING (red box) are conducted to identify anomalous patterns that occurred in Brazilian state capitals from 2020 to 2023, drawing comparisons to the events occurred during the whole COVID-19 pandemic period between 2020 and 2022. Detection of possible outbreaks in original and synthetic data and comparison with EARS results are evaluated using statistical metrics.
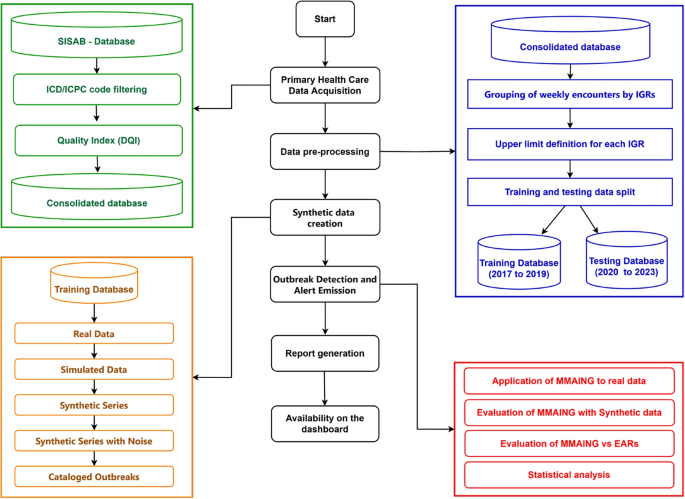
Simplified MMAING workflow with main steps in the central column and details in the lateral ones. The green box represents the initial stages related to data acquisition from Brazil’s National Primary Health Information System (SISAB), filtering of diseases related to URTI using the International Classification of Diseases (ICD-10) and the International Classification of Primary Health Care (ICPC-2) codes, and Data Quality Index (DQI) evaluation. The blue box identifies the pre-processing stages, such as grouping the data at municipality level by Imediate Geographic Regions (IGR), calculation of the upper limit, and data splitting. The orange box indicates the stages for generating and cataloging the synthetic series. The red box describes the stages for Outbreak Detection, EWS Emission, and the comparison of MMAING with EARS on real and synthetic data. The reports and dashboard access are currently of exclusive use by the responsible health teams at municipal, state and national level
Data sources
We used data from Brazil’s National Primary Health Information System (SISAB – Portuguese acronym for Sistema de Informação em Saúde para a Atenção Básica), which contain data on all encounters from publicly funded primary health care (PHC) facilities. The public healthcare in Brazil is universal and covers the entire population, and around 75% of the population only use the public healthcare [24]. Each encounter in PHC healthcare facilities is coded using International Classification of Diseases (ICD-10) and International Classification of Primary Health Care (ICPC-2). The original data are organized by municipality (identified by name and code), year, epidemiological week. This study, covering data for the period 2017–2023, is based on a data subset that is obtained by extracting only those entries with codes related to 50 specific upper respiratory tract infection (URTI) conditions. After this, the data is organized into time series, where each data point corresponds to the weekly count PHC visits in that municipality related to URTIs. The detailed list of the 50 ICD-10 and ICAP-2 codes can be found in the Supplementary Material (MS) Table A1. Before being used in the analyses, the data undergoes a quality test, which takes into account the completeness, timeliness and consistency of the records. This test assigns to each municipality its weekly Data Quality Index (DQI) indicating whether these criteria were satisfied or not. Analyses are performed only for those pieces of data with a positive DQI evaluation [25].
Data from the 5,570 Brazilian municipalities were grouped into 510 Immediate Geographic Regions (IGRs) for a precise analysis of the risk of epidemic outbreaks. As defined by the Brazilian Institute of Geography and Statistics (IBGE), an IGR is a group of municipalities that have, as their primary references, an urban network and a local urban center where the nearby population seeks goods, services, and work. We used the URTI time series of 27 IGRs corresponding to capital cities, covering 41% of the country’s inhabitants, offering a broad and diversified view of the evolution of respiratory syndromes during the epidemiological weeks of the period under study.
Study design
We combined four unsupervised methods (ISF, LOF, OCSVM, COPOD) commonly used for anomaly detection with the deterministic NGM that accounts for dynamic causes, underlying formation, and outbreak propagation. These five methods were aggregated into an ensemble (MMAING), representing an integrative methodology to identify emerging epidemic patterns through the detection of EWS. The association between series anomalies and EWS requires that the former be restricted to upward trends. The final indication of pandemic risk (yes/no) is based on a voting system that takes into account the equally weighted yes/no EWS indications provided by each of the methods individually. The final prediction is the option receiving the mojority of votes, which can unambiguously be applied for an odd number of methods [26, 27].
To improve the efficiency in detecting EWS among different models, where the identification of detected points can be counterproductive, we propose a strategy consisting of establishing a time-dependent upper limit \(\epsilon (x(t))\) based on confidence intervals [28], and assigning a positive risk of outbreak only to those values of t exceeding such threshold, i.e., \(x(t)> \epsilon (x(t))\). Mathematically, this threshold is defined as
$$\begin{aligned} \epsilon (x) = \bar{x} + \frac{z \sigma }{\sqrt{n}}, \end{aligned}$$
(1)
where \(\bar{x}, z,\,\sigma\) and n denote the sample mean, the critical value for a given confidence level, which we assumed to be z=1.96 to warrant a 95% of confidence interval. The sample standard deviation, and the sample size, respectively. This threshold is crucial for identifying potential EWS, but its effectiveness depends on the sampling strategy adopted. We applied two strategies:
-
A moving window scheme averages the data from a recent past of w weeks, removing short-term fluctuations. This approach proves to be extremely useful when there is a need to monitor recent trends in the data set [29].
-
Seasonally adjusted regimes considering cyclical patterns or trends from previous years. This choice is essential for events impacted by external variables, such as climate change, holidays, and other periodic events.
Our methodology considered both strategies, with \(n=5\) for the moving window scheme and references to the same epidemiological weeks from 2017 to 2019 for the seasonal scheme. These strategies facilitate the detection of increased patterns at specific time intervals, ensuring that only EWS points exceeding the threshold value are considered while improving precision by avoiding false positives. The indication of a potential outbreak is followed by validation against already established syndromic surveillance methods, such as EARS and synthetic data.
Synthetic data
To evaluate the models used in this study, we created 27 synthetic series based on real data from each of the 27 IGRs of Brazilian capital cities, as illustrated in Fig. 2. We introduced noise into the generated series to simulate periods of abnormal behavior, which we defined as outbreaks, and recorded these disturbed intervals to identify and catalog these occurrences, similarly to the approach reported by Neill [30]. This procedure allowed the evaluation of the models using statistical metrics. To do so, we used real data recorded between 2017 and 2019, before the COVID-19 pandemic, and carried out the following procedures:
Real series smoothing
Let \(X_t\), where t is a discrete variable ranging from \(t_1=1\) to \(t_N=N\), be a time series representing a value recorded for an epidemiological week. The moving average \(M_i\) at each time \(t_i\) is obtained by convolving the data \(X_{t_i}\equiv X_i\) with a moving window of any width \(1
$$\begin{aligned} M_i = \frac{1}{8} \sum \limits _{t = i-3}^{i + 4} X_i. \end{aligned}$$
(2)
Synthetic data generation
The moving average converts the original series of integer number is into a series of continuous rational numbers. Therefore, for each time \(t_i\), a normal continuous distribution \(\mathcal {N }(\mu , \sigma )\) was used to generated synthetic data \(\bar{X_i} \sim \mathcal {N }(\mu , \sigma )\) as the average of (\(n = 200\)) simulations, whereby
$$\begin{aligned} \mathcal {N}(\mu , \sigma ) = \frac{1}{\sigma \sqrt{2\pi }} \exp \left[ -\frac{1}{2}\left( \frac{x_t-\mu }{\sigma }\right) ^2\right] \end{aligned}$$
(3)
The mean \(\mu\) of each distribution was set to that of the smoothed series \(M_i\) multiplied by a factor \(F_M\) ranging from 0.8 to 1.2. The standard deviation \(\sigma\) was taken as that of the real series, multiplied by a factor \(F_D\) ranging from 0.2 to 0.8. This procedure allowed the generation of a data series following a distribution similar to the original series with superimposed small changes.
Addition of random noise
To test the ability of MMAING and other methods to detect specific events, we generated the series \(Z_i\) to simulate the emergence of outbreak periods by adding random fluctuations to the series \(\bar{X_i}\). Each of the \(K \in [1,6]\) simulated outbreaks, which may last from 4 to 10 weeks, consists of a set of \(4 \le P_k \le 10\) consecutive points, where \(P_k\) is randomly chosen from a uniform distribution U(4, 10). Its starting time \(t_k\) is also randomly chosen from \(U(1,N-10)\). This procedure does not exclude the possibility that a specific time \(t_j\) be included in two or more sets \(P_k\). This way, after starting with the series \(\bar{X_i}\), a total number \(L=\sum _{k=1}^{K}P_k\) of interventions is required to obtain \(Z_i\). Each intervention \(\ell\) in the interval [1, L] amounts to evaluating \(\tilde{\epsilon _{\ell }}\) to be added to a well defined \(\bar{X_{\ell }}\), in such a way that
$$\begin{aligned} Z_{\ell } = \bar{X_{\ell }}+\tilde{\epsilon _{\ell }}. \end{aligned}$$
(4)
For a given series \(\bar{X}_i\), the value of each \(\tilde{\epsilon _{\ell }}\) is obtained by a fixed procedure, which is independent of the value of \(\ell\) and of the interval \(P_k\) where \(\bar{X_{\ell }}\) is located. It requires the random choice of two parameters from two previously evaluated sets, namely the amplitude \(\textbf{A}=\{A_1, A_2, …, A_{10}\}\) and frequency \(\textbf{f}=\{f_1, f_2,…,f_8\}\), where \(A_j \sim U[50,B]\), with B as an integer number defined as \(B=(max(\bar{X}_-min(\bar{X}))/2\), so that the set \(\textbf{A}\) depends on the specific series, and \(f_j \sim U[0,1]\) does not explicitly depends on the series \(\bar{X}_i\). Finally, with the help of \(\textbf{A}\) and \(\textbf{f}\), we define
$$\begin{aligned} \tilde{\epsilon _{\ell }} = A_{\ell } |(\sin (2 \pi f_{\ell }(t_k – t_{\ell } ) + \frac{1}{10^4}|, \end{aligned}$$
(5)
where the inclusion of a small non-zero constant in the definition of \(\tilde{\epsilon }_{\ell }\) seeks to realistically reproduce the initial behavior of an outbreak, playing an essential role to simulate an actual event.
Cataloguing of abnormal occurrences introduced by addition of noise
To verify whether the models would be able to detect new variations and allow an analysis through statistical metrics, we first identified the abnormal signals presented in the series \(\bar{X_i}\) without noise, which may have originated from the preservation of the existing trend in the original series. Then, we recorded the perturbed periods by adding noise (\(\tilde{\epsilon }\)) to create a reference dataset, identifying the start and end points of the outbreak, which will be the detection target.
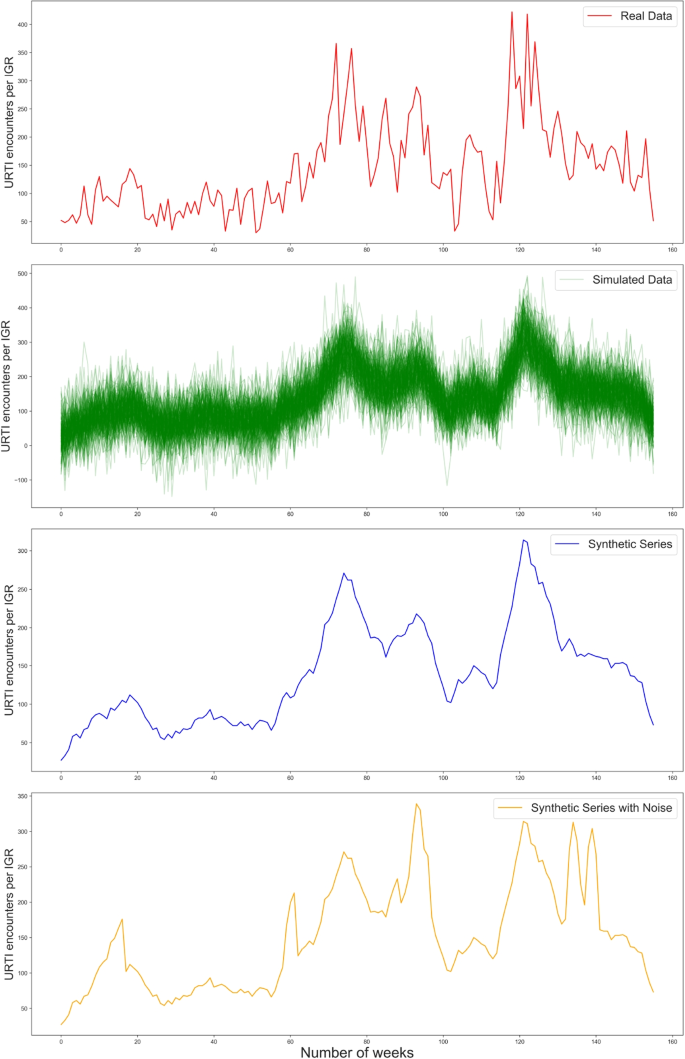
Process of synthetic data generation: Red – real data from an IGR; Green – set of 200 simulated series; Blue – synthetic time series by averaging over green curves; Orange – synthetic time series with superimposed noise
MMAING ensemble
In machine learning (ML), an ensemble combines multiple models aiming to achieve higher precision and robustness while reducing variation and bias compared to single, isolated models. Among the existing approaches, hard voting [27, 31] stands out as one of the simplest and most effective. This specific technique consists of training a plurality of different models and subsequently employing a model, which in the context of this work is called MMAING, to integrate the results of the base models. The original data set is used to train the base models, whose results serve as input for MMAING, culminating in the decision about issuing an EWS or not, as illustrated in Fig. 1.
MMAING’s innovation is materialized through a previously unexplored combination of four ML models and one NGM model and a selective strategy that chooses signals from three of them based on a majority voting system that guarantees an equal contribution of each model. Therefore, MMAING not only simplifies the interpretation of results through an integrative voting process but also reinforces the reliability of detections, establishing a robust consensus among its methods.
Machine learning methods
Unsupervised machine learning models are commonly applied to anomaly detection over time series data, as they can learn patterns from the data and thus spot those points that do not fit any patterns as anomalies [32, 33]. MMAING incorporates the four ML methods that will be detailed in the sequence. They perform better in detecting anomalies as compared to other unsupervised methods targeted to predict continuous outcomes in regression problems, like OLS, LASSO, SVR, Boosting, etc.
Isolation Forest (ISF):
a tree-based algorithm that uses the concept of isolation to calculate an abnormality score for each point and employs a recursive partitioning process to isolate outliers. ISF creates a random forest, where each decision tree randomly selects a feature and chooses a threshold value within the minimum and maximum range of that feature to split the data. This process is repeated until all the data points are isolated. Anomalies are identified based on a score value obtained by the average length of paths in the forest trees divided by a normalization factor. Abnormal data points tend to be isolated more quickly, resulting in shorter paths from the root compared to standard data points [17, 18].
Local Outlier Factor (LOF):
an algorithm designed to identify outliers in diverse datasets, including time series. LOF works by evaluating the local density (i.e., how tightly packed) of data points compared to the density of their neighbors. Consequently, a point surrounded by many nearby points will have a high local density, while an isolated point will exhibit a low local density. If the local density of a point is significantly lower relative to its neighbors, it is classified as an outlier [19].
One-Class Support Vector Machine (OCSVM):
an algorithm aiming to find an optimal hyperplane that maximizes the separation between examples of different classes in a high-dimensional space. This process involves projecting the input data into a high-dimensional space via a kernel function and creating a hyperplane to determine whether a new observation is within the hyperplane (not an anomaly) or outside it (an anomaly) [20].
Copula-based outlier detection (COPOD):
it represents a significant innovation in the field of multivariate outlier detection, moving away from conventional cluster-centric approaches. Unlike these approaches, COPOD is based on copulas [34], a mathematical function to model dependencies (correlation) between variables. Detecting outliers with COPOD involves a three-step process: i) calculating empirical cumulative distribution functions based on the data, ii) generating the empirical copula function from these functions, and iii) using the empirical copula to estimate the tail probabilities and quantifying these probabilities as data anomaly scores [21]. Tail probabilities correspond to the probability of a data point falling at the extremes of distributions, which are the least frequent areas and, therefore, most susceptible to outliers. After estimating the tail probabilities, COPOD transforms them into anomaly scores representing the degree of abnormality of each point, and those points with high abnormality degrees are flagged as potential outliers.
Next Generation method
The Next Generation Method (NGM) provides reliable information on the evolution of epidemic outbreaks, including the reproduction number \(R_0\), which measures the average number of new infections caused by an infected individual at the beginning of an epidemic process when the whole population has no protection to that infection [35, 36]. Recently, that procedure was generalized in terms of the generation time interval distribution [22, 37], for any value of \(t>0\). Its implementation requires the time series of confirmed cases B(t) in a given population, from which one may infer the number of individuals who become infected and the number of others to whom they can transmit the pathogens. R(t) estimates are based on classical compartmental models (SIR, SEIR, and more complex ones) [22, 23, 38] or on approaches using data of confirmed cases without further consideration of a model dynamics [39,40,41,42].
As in other problems of population dynamics, the estimation of R(t) needs the idea of generation time, so that the variables of the dynamical system have to depend on both time t and age \(\tau\). For epidemiological systems, \(\tau\) means the age of infection [43], and as a consequence R(t) represents the average number of people that an individual infected at time t can infect during his entire infectious period \(\tau\). The implementation of NGM to accurately describe the spread of the agent in the population requires the accurate identification of each individual who becomes infected and the number of contacts with other individuals to whom the infected person can transmit the pathogens.
Using a generation interval distribution, also called generation time distribution, \(g(\tau )\), which considers the time that an individual who visited a PHC unit, if infected, takes to generate a new infection, we define R(t) as:
$$\begin{aligned} R(t) = \frac{a(t)}{\int _{0}^{\infty } a(t-\tau ) g(\tau ) d\tau } \end{aligned}$$
(6)
where, a(t) is the number of visits and \(g(\tau )\) is the normalized generation time distribution.
According to [23], the generation time interval distribution \(g(\tau )\) for the SEIR compartmental model is given by:
$$\begin{aligned} g(\tau ) = \frac{\epsilon e^{-\kappa \tau } + \frac{\kappa }{\gamma -\kappa } [e^{-\kappa \tau } – e^{ -\gamma \tau }]}{\frac{\epsilon }{\kappa }+\frac{1}{\gamma }} \end{aligned}$$
(7)
for which the parameters \(\epsilon\), \(\kappa\) and \(\gamma\) represent, respectively, factor of pre-symptomatic infection, inverse of latency period, and removal rate. In this study, given the lack of knowledge about the disease with respiratory symptoms and therefore its dynamics, we defined \(\kappa\) and \(\gamma\) as equal (\(\kappa = \gamma + \delta\) with \(\delta \rightarrow 0\)), and \(\epsilon = 0\) as in the simple version of SEIR model. Thus, \(g(\tau )\) reduces to:
$$\begin{aligned} g(\tau ) = e^{-\gamma \tau } \gamma ^2 \tau \end{aligned}$$
(8)
To be conveniently used, this form of the distribution \(g(\tau )\) needs to be converted to the corresponding version valid for discrete time and then normalized [44]. For this, we will discretize this distribution by initially considering a geometric progression associated with SEIR model. Replacing \(\tau\) by \((n-1)\) in (8), the general term of the geometric progression is given by:
$$\begin{aligned} g(n) = (n-1) g_1 q^{(n-1)}, \end{aligned}$$
(9)
where \(q = e^{-\gamma }\), \(t = 0\) corresponds to \(n = 1\). Based on the sum S(m) of the first m terms of the function g(n), the first term \(g_1\) (the normalization factor) is obtained (for the intermediate steps of the calculation see Appendix B.1), leading to:
$$\begin{aligned} g_1 = \frac{(1-q)^2}{q[1+(m-1)q^m – mq^{m-1}]} \end{aligned}$$
(10)
Therefore, we are applying the method based on the Next Generation Matrix technique (NGM) discretized and normalized in terms of a geometric progression, and, as in [23] we take into account the dynamics through \(g(\tau )\) and the data through a(t).
In order to promote a more realistic potential detection, MMAING adds a new perspective to this metric by including the estimation of \(\hat{R}(t)\), which is defined in a similar way as R(t), with the difference that it now depends on the weekly incidence of registered encounters at PHC posts. It corresponds to an extension to the approach developed in [23], which amounts to using the series of URTI-related PHC encounters to evaluate \(\hat{R}(t)\), an analogous of usual R(t) that might assess the risk of epidemic outbreaks. If we use the same expressions for evaluating R(t) in [23] as well as similar transmission dynamics based on the SEIR model, some key mathematical properties of R(t) are also expected to hold for \(\hat{R}(t)\), e.g. an increase or decrease of its values when the number of encounters increases or decreases.
By contrast, the clear-cut epidemiological meaning of R(t) [44, 45] (whether smaller, equal or larger than 1) obtained from a series of confirmed cases cannot be transferred to \(\hat{R}(t)\) in a straightforward way, but it becomes necessary to calibrate the threshold value of \(\hat{R}(t)\) that better measures the actual outbreak risk. Because the number of confirmed infection cases is usually smaller than the number of encounters, it is expected that \(\bar{R}\), a threshold value of \(\hat{R}(t)\) defining actual epidemic risk, will be larger than 1. Indeed, our estimates have shown that \(\langle \hat{R}(t)\rangle\), the average of \(\hat{R}(t)\) taken over values of \(\hat{R}(t)>1\) for the national PHC time series is \(\sim 1.24\). Additionally, the analyses of the corresponding series for each Brazilian state consistently indicate that \(1.2< \langle \hat{R}(t)\rangle < 1.3\). Therefore, we established the criterion for outbreak risk by the condition \(\hat{R}(t)> 1.25 \equiv \bar{R}\). According to that criterion, the NGM is applied carrying to the ensemble information from both dynamical process and on the data.
A somewhat similar analysis was used in a study on the transmissivity of Ebola virus disease based on confirmed cases, where it was observed that a suitable threshold value \(\bar{R}\) for the usual R(t), set in terms of its past median, could forecast spreading trends within 1–2 weeks [46]. These results were confirmed one year later in a similar study using COVID-19 data [10].
Early Aberration Reporting System (EARS)
EARS [6] is available in its three variants – C1, C2, and C3 – mainly used to monitor weekly syndromic counts. These methods are helpful when limited baseline data is available. The variants are based on the Shewhart procedure that uses a moving sample mean and a sample standard deviation to standardize each observation [47]. By default, variant C1 calculates the sample mean and standard deviation using data from the seven weeks before the current observation; C2 is similar but considers a two-week delay; and C3 combines the results obtained with C2 for the current and two previous weeks, as detailed discussed in [6, 47]. In this work, however, we extended the period to eight weeks to better adjust the data. EWS for the different variants are produced when the corresponding statistics C1 or C2 exceed three sample standard deviations above the sample mean or if C3 exceeds two sample standard deviations above the sample mean [6, 47].
MMAING configuration
As mentioned earlier, MMAING is an ensemble of models that utilizes a voting system to detect EWS linked to potential outbreaks in syndromic time-series data. Data recorded between 2017 and 2019 was used for training due to the stability observed in the number of encounters throughout epidemiological weeks to ensure sensitivity to abnormal changes. Subsequently, we applied these models to real datasets recorded between 2020 and 2023, aiming to empirically analyze the anticipation of outbreak periods during the COVID-19 pandemic. Additionally, we evaluated the performance of the proposed ensemble with synthetic data.
Furthermore, to assess MMAING’s adaptability and versatility in scenarios that may demand moderate or high rigor (i.e., larger or smaller number of false positive EWS), we adopted two distinct configurations, which distinguish themselves by the values of a few specific parameters in each method. The parameter values adopted in each configuration, named “balanced” (BLCf) and “strict” (STCf), are indicated in Table 1. In both cases, the adopted configurations were used to run all ML models were based on a total of 810 synthetic series obtained from 30 independent series for each of the actual data sets for the 27 IGRs. BLCf settings may decrease precision and increase sensitivity, resulting in more EWS and consequently increasing the number of false positives. On the other hand, the STCf aims to enhance precision and reduce the false positive rate, which may result in failing to issue true EWS.
BLCf uses parameters that seek a compromise between sensitivity and specificity. For example, for the ISF and LOF methods, the number of estimators (\(n_{est}\)) and neighbors (\(n_{nei}\)) are set to 500, while the contamination \(\mathcal {C}\) is set to 0.4, indicating that we expect about 40% of anomalous points. The strict configuration means a reduction in the number of ISF trees and the number of LOF neighbors to 400 and 300, respectively, as well as a reduction in contamination to 0.3.
The OCSVM method uses a higher \(\nu\) value in BLCf (0.8) compared to STCf (0.5), indicating greater flexibility in class separation. The use of the Radial Basis Function (RBF) kernel and the value of \(\gamma\) are kept consistent between configurations, suggesting that the shape of the decision boundary and the complexity of the model are considered adequate in both cases. COPOD and NGM also present different parameter values between configurations. COPOD maintains consistency with ISF and LOF concerning contamination, while NGM adjusts the threshold \(\bar{R}\) to reflect the desired stringency of detection.
In outbreak detection contexts, these configurations can be adapted with specific objectives to monitor public health data. While the sensitivity measure is important if a clear and consistent EWS is needed, in scenarios where surges occur quite frequently, precision (or positive predictive value) measures the probability of an EWS being true, particularly when surges do not occur frequently. For each of these measures, the user may want to define settings for the algorithms and/or prioritize which measures are most important for their surveillance needs [48].
Training and validation
To assess MMAING’s effectiveness, we adopted three distinct approaches. The first involved using PHC data to conduct an empirical analysis, comparing the outbreak detection results obtained during the COVID-19 pandemic with the official reports from the Brazilian government [49], which detail the different waves of the pandemic. The second approach aimed to evaluate the detection capability of MMAING under two distinct configurations using categorized synthetic data. Finally, the third approach used real and synthetic data to compare MMAING and EARS models. To achieve this, we employed statistical metrics such as Probability of Detection (POD), Sensitivity, Specificity, Positive Predictive Value (PPV), and F1 [12, 50], under the following assumptions:
-
POD evaluation: For each scenario and each period of the current week, if an EWS is generated at least once between the start and end of an outbreak, the outbreak is considered detected [8, 48]. POD is an event-based sensitivity (i.e., the entire outbreak interval is counted as a single observation for the sensitivity measurement), thus corresponding to the proportion of outbreaks detected with the total number of synthetic replicas.
-
For sensitivity analysis, True Positive (TP), True Negative (TN), False Negative (FN), and False Positive (FP) detected events are defined, respectively, as weeks with a surge and issued a warning, weeks with no surges nor warning, weeks with a surge but no warning, and weeks with no surge but an issued warning.
-
All metrics are defined as usual: \(Sensitivity (Se) = TP/(TP+FN)\), \(Specificity = TN/(TN+FP)\); Positive Predictive Value: \(PPV = TP/(TP+FP)\), and F1 measurement, defined as the harmonic mean of sensitivity and PPV: \(F1 = 2 \times (Se \times PPV)/(Se + PPV)\). Finally, the average reliability is defined as the average of the previous five metrics.
Codes
All calculations were perfomed using the Python language. For the ML methods, we used the Scikit-learn library, including the Isolation Forest, One Class SVM and Local Outlier Factor algorithms. We also used the PyOD library for COPOD and sklearn.metrics for the analyses. The NGM code has been created by the authors. All codes are available at
link